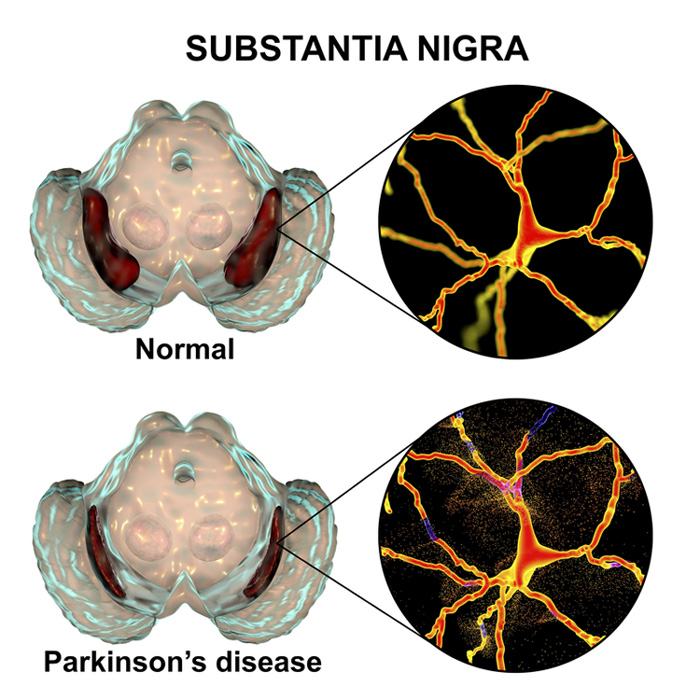
10th May 2023 AI detects Parkinson's disease with 96% accuracy An algorithm to detect Parkinson's disease, years before the onset of symptoms, has been demonstrated by the University of New South Wales and Boston University. CRANK-MS can achieve 96% accuracy, using neural networks to analyse biomarkers in bodily fluids.
No blood test is currently available to identify the risk of non-genetic Parkinson's disease. But that may soon change if a new machine-learning tool is approved and deployed in medical settings. Scientists from the University of New South Wales (UNSW), in a collaboration with Boston University, have used artificial intelligence to detect this chronic, degenerative disorder years before the onset of symptoms. In their study, published this week in ACS Central Science, the team combined a new algorithm with neural networks to analyse biomarkers in patients' bodily fluids. The research focused on 39 people who developed Parkinson's up to 15 years after the samples were taken. The AI looked through datasets containing extensive information about metabolites – chemical compounds that the body creates when breaking down food, drugs, or chemicals.
After comparing these metabolites to those of 39 matched control patients – people in the same study who didn't go on to develop Parkinson's – the team were able to identify unique combinations of metabolites that could prevent or potentially be early warning signs for Parkinson's. Triterpenoids, for example, appeared in lower concentrations in the blood of those who later developed Parkinson's compared to those who did not. These are a neuroprotectant that regulate oxidative stress and are commonly found in foods such as apples, olives, and tomatoes. A future study could examine whether eating these foods can naturally protect against developing Parkinson's disease. Also of note is that polyfluorinated alkyl substances (PFAS) appeared in people who went on to develop Parkinson's, which could be linked to industrial chemical exposure. The algorithm, called CRANK-MS (Classification and Ranking Analysis using Neural network generates Knowledge from Mass Spectrometry) is user-friendly and can generate results in less than 10 minutes on a conventional laptop. Its creators have made it publicly available to any other researchers who would like to use machine learning for disease diagnosis using metabolomics data. Validation studies are needed, using larger cohorts from around the world. But in the limited cohort for this study, the results are promising, with CRANK-MS able to analyse chemicals found in blood to detect Parkinson's disease with up to 96% accuracy.
"The most common method of analysing metabolomics data is through statistical approaches," said Diana Zhang, Scientia PhD Scholar at UNSW. "So to figure out which metabolites are more significant for the disease versus control groups, researchers usually look at correlations involving specific molecules. But here we take into account that metabolites can have associations with many other metabolites – which is where the machine learning comes in. With hundreds to thousands of metabolites, we've used computational power to understand what's going on." In addition to looking at combinations of metabolites, the researchers used an unedited list of data, as explained by W. Alexander Donald, study co-author and Associate Professor in the School of Chemistry at UNSW. "Typically, researchers using machine learning to examine correlations between metabolites and disease reduce the number of chemical features first, before they feed it into the algorithm," said Donald. "But here we feed all the information into CRANK-MS without any data reduction right at the start. And from that, we can get the model prediction and identify which metabolites are driving the prediction the most, all in one step. It means that if there are metabolites which may potentially have been missed using conventional approaches, we can now pick those up." "The application of CRANK-MS to detect Parkinson's is just one example of how AI can improve the way we diagnose and monitor diseases," added Ms Zhang. "What's exciting is that CRANK-MS can be readily applied to other diseases to identify new biomarkers of interest."
Comments »
If you enjoyed this article, please consider sharing it:
|