22nd February 2020 3D atomic-scale image could lead to coronavirus vaccine Using cryogenic electron microscopy – a Nobel Prize-winning technique – scientists have produced an atomic-level image of an essential protein required to access cells by SARS-CoV-2. This 3D image could help in more quickly finding a treatment for the viral infection that is spreading around the world.
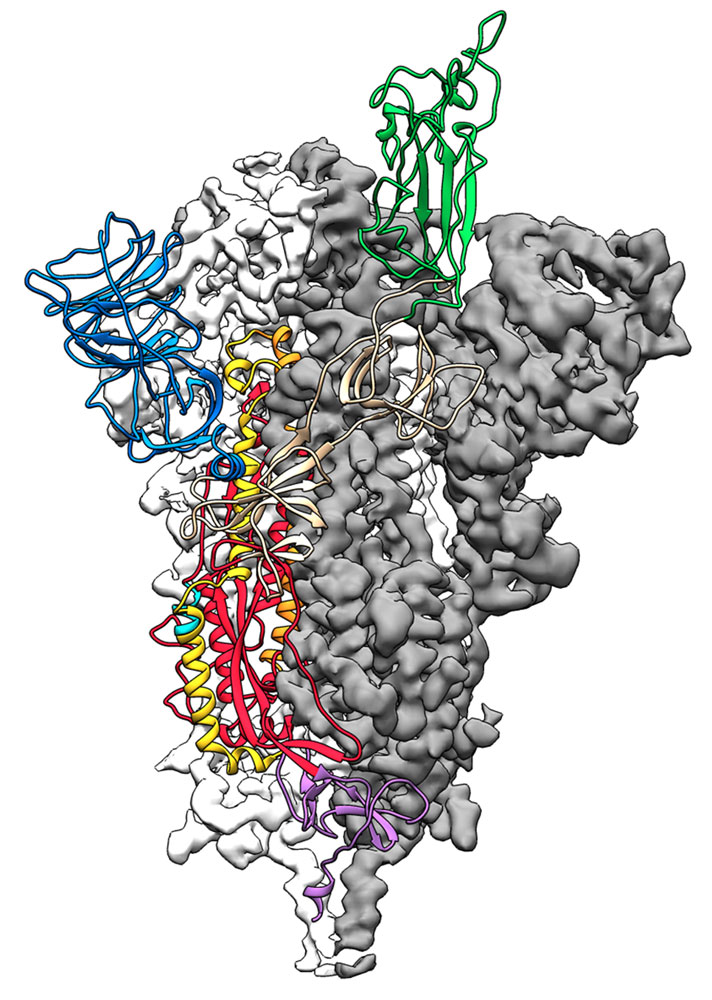
Researchers from The University of Texas at Austin and the National Institutes of Health have made a critical breakthrough toward developing a vaccine for the 2019 novel coronavirus. Their work has produced the first 3D atomic-scale map of the "spike protein" – the part of the virus that attaches to and then infects human cells – which could allow medical facilities around the world to more quickly develop vaccines and antiviral drugs. The same team is already working on a potential vaccine candidate stemming from the research. Jason McLellan, associate professor at UT Austin who led the research, and his colleagues have spent many years studying other coronaviruses, including SARS-CoV and MERS-CoV. They had already developed ways of locking coronavirus spike proteins into a shape that made them easier to analyse and this experience gave them an advantage over other research teams. "As soon as we knew this was a coronavirus, we felt we had to jump at it, because we could be one of the first ones to get this structure. We knew exactly what mutations to put into this, because we've already shown these mutations work for a bunch of other coronaviruses," explains McLellan. The bulk of the research was carried out by the study's co-first authors, postdoctoral research fellow Nianshuang Wang (seen here on the right) and PhD student Daniel Wrapp (to his left), both at UT Austin.
Just two weeks after receiving the genome sequence of the virus from Chinese researchers, the team had designed and produced the samples of their stabilised spike protein. It took about 12 more days to fully reproduce the molecular structure of the spike protein and then submit a manuscript to the journal Science, which expedited its peer review process. The many steps involved in this process would typically take months to accomplish. Critical to their success was a state-of-the-art technology known as cryogenic electron microscopy (cryo-EM) used in the university's new Sauer Laboratory for Structural Biology. Cryo-EM allows researchers to make atomic-scale 3D models of cellular structures, molecules and viruses. The developers of Cryo-EM won the Nobel Prize in Chemistry 2017 "for developing cryo-electron microscopy for the high-resolution structure determination of biomolecules in solution." The process involves quickly "freezing" a molecule in place and then bombarding it with electrons to create over 3,000 two-dimensional projections, which are then combined to create a 3D reconstruction of the protein. This was made possible thanks to a recently installed $3 million electron microscope. "We ended up being the first ones in part due to the infrastructure at the Sauer Lab. It highlights the importance of funding basic research facilities," explains McLellan. The molecule the team produced, and for which they obtained a 3D structure, represents only the extracellular portion of the spike protein – but is enough to elicit an immune response in people, and thus serve as a vaccine. Next, McLellan's team plans to use their molecule to pursue another line of attack against the virus that causes COVID-19, using the molecule as a "probe" to isolate naturally produced antibodies from patients infected with the novel coronavirus who successfully recovered. In sufficient quantities, these antibodies could help treat a coronavirus infection soon after exposure. For example, the antibodies could protect soldiers or health care workers sent into an area with high infection rates on too short notice for the immunity from a vaccine to take effect. The Science paper was published on 19th February and is available here.
Comments »
If you enjoyed this article, please consider sharing it:
|